So here is a recap of the previous post:
A simple problem with hierarchical Pitman-Yor processes (PYPs) or Dirichlet processes (DP) is where you have a hierarchy of probability vectors and data only at the leaf nodes. The task is to estimate the probabilities at the leaf nodes. The hierarchy is used to allow sharing across children …
We have a tree of nodes which have probability vectors. The formula used estimates a probability for node 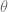 from its parent
from its parent 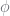 as follows:
as follows:
 (1)
(1)
In this, the pair  are the parameters of the PYP called discount and concentration respectively, where
are the parameters of the PYP called discount and concentration respectively, where  and
and  . The
. The  are magical auxiliary (latent) counts that make the PYP samplers work efficiently. They are, importantly, constrained by their corresponding counts, and the counts are computed up from the leaves up (using
are magical auxiliary (latent) counts that make the PYP samplers work efficiently. They are, importantly, constrained by their corresponding counts, and the counts are computed up from the leaves up (using  to denote
to denote 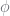 are the children of
are the children of  ). The
). The 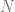 s and
s and  s are totals. These relationships are summarised in the table below.
s are totals. These relationships are summarised in the table below.
| constraint |
 |
| constraint |
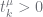 whenever whenever 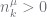 |
| propagating counts |
 |
| total |
 |
| total |
 |
It is important to note here how the counts propagate upwards. In some smoothing algorithms, propagation is  . This is not the case with us. Here the non-leaf nodes represent priors on the leaf nodes, they do not represent alternative models. PYP theory tells us how to make inference about these priors.
. This is not the case with us. Here the non-leaf nodes represent priors on the leaf nodes, they do not represent alternative models. PYP theory tells us how to make inference about these priors.
Now the theory of this configuration appears in many places but Marcus Hutter and I review it in Section 8 of an ArXiv paper, and its worked through for this case in my recent tutorial in the section “Working the n-gram Model”. The posterior distribution for the case with data at the leaf nodes is given by:
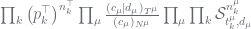 (2)
(2)
There are a lot of new terms here so let us review them:
 |
is the Pochhammer symbol (a generalised version is used, and a different notation) which is a form of rising factorial so its given by  ; can be computed using Gamma functions ; can be computed using Gamma functions |
 |
a related rising factorial given by 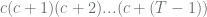 so it corresponds to so it corresponds to  |
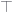 |
the root node of the hierarchy |
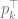 |
the initial/prior probability of the k-th outcome at the root |
 |
a generalised Stirling number of the second kind for discount  and counts and counts  |
Background theory for the Stirling number is reviewed in Section 5.2 of the ArXiv paper, and developed many decades ago by mathematicians and statisticians. We have C and Java libraries for this, for instance the C version is on MLOSS so its O(1) to evaluate, usually.
Now to turn this into a Gibbs sampler, we have to isolate the terms for each  individually. Let
individually. Let 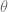 be the parent of
be the parent of  . The constraints are:
. The constraints are:
| constraint |
 |
| constraint |
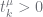 whenever whenever 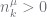 |
| propagating counts |
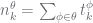 |
| propagating counts |
 |
The propagating constraints do not apply when  is the root, and in this case, where $\vec{p}^\top$ is given,
is the root, and in this case, where $\vec{p}^\top$ is given,  does not exist. For
does not exist. For 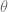 the parent of
the parent of  , the constraints can be rearranged to become:
, the constraints can be rearranged to become:
The terms in Equation (2) involving  when
when  is not the root are:
is not the root are:
 (3)
(3)
where note that  appears inside
appears inside 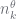 ,
,  and
and  . When
. When  is the root, this becomes
is the root, this becomes
 (4)
(4)
The Gibbs sampler for the hierarchical model becomes as follows:
Gibbs sampler
For each node  do:
do:
for each outcome  do:
do:
sample  according to Equation (3) or (4) satisfying constraints in the last table above.
according to Equation (3) or (4) satisfying constraints in the last table above.
endfor
endfor
As noted by Teh, this can result in a very large range for  when
when 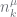 is large. We overcome this by constraining
is large. We overcome this by constraining  to be within plus or minus 10 of its last value, an approximation good enough for large
to be within plus or minus 10 of its last value, an approximation good enough for large 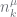 .
.
Moreover, we can initialise  to reasonable values by propagating up using the expected value formula from Section 5.3 of the ArXiv paper. We only bother doing this when
to reasonable values by propagating up using the expected value formula from Section 5.3 of the ArXiv paper. We only bother doing this when  . The resultant initialisation procedure works in levels: estimate
. The resultant initialisation procedure works in levels: estimate  at the leaves, propagate these to the
at the leaves, propagate these to the 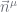 at the next level, estimate
at the next level, estimate  at this next level, propagate these up, etc.
at this next level, propagate these up, etc.
Initialisation
for each level, starting at the bottom do:
for each node  at the current level do:
at the current level do:
for each outcome  do:
do:
if 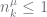 then
then

elseif  then # the PYP case
then # the PYP case
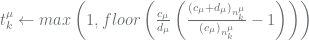
else # the DP case

#  is the digamma function
is the digamma function
endif
endfor
endfor
for each node  directly above current level do:
directly above current level do:

endfor
endfor
This represents a moderately efficient sampler for the hierarchical model. To make this work efficiently you need to:
- set up the data structures so the various counts, totals and parent nodes can be traversed and accessed quickly;
- set up the sampling so leaf nodes and parents where
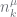 is guaranteed to be 0 or 1 can be efficiently ignored;
is guaranteed to be 0 or 1 can be efficiently ignored;
- but more importantly, we also need to sample the discounts and concentrations.
Note sampling the discounts and concentrations is the most expensive part of the operation because each time you change them the counts  also need to be resampled. When the parameters are fixed, convergence is fast. We will look at this final sampling task next.
also need to be resampled. When the parameters are fixed, convergence is fast. We will look at this final sampling task next.
samples, Mark’s first plot shows the expected number of clusters,
, that one would get with a DP using concentration parameter
. The thing to notice is that the number of clusters is moderately well determined by the concentration parameter. In fact the mean (expected value) of
is given by:
is the digamma function; for details see the ArXiv report by Marcus Hutter and I, Section 5.3. Morever, the standard deviation of
is approximately the square root of the mean (for larger concentrations).
, on the concentration, we now smear out the previous plots. So now we show plots for different values of
. As you can see, these plots have a much higher variance, which is what you want. With a given
, you are still determining the broad range of the number of clusters, but you have a lot more latitude.
(usually) by sampling it. If we use Chinese restaurant processes, there is a simple auxiliary variable sampling formula for the concentration (presented in the original HDP paper by Teh et al.). If we use our blocked table indicator sampling, the posterior on the concentration is log concave so we can use either slice sampling or adaptive rejection sampling (ARS). The implementation is moderately simple, and it works well. However, it does mean now your algorithm will expend more time as it has to try and find the right concentration as well.


